Friday, March 26, 2010
Does a forty thousand year old finger point to another human species?
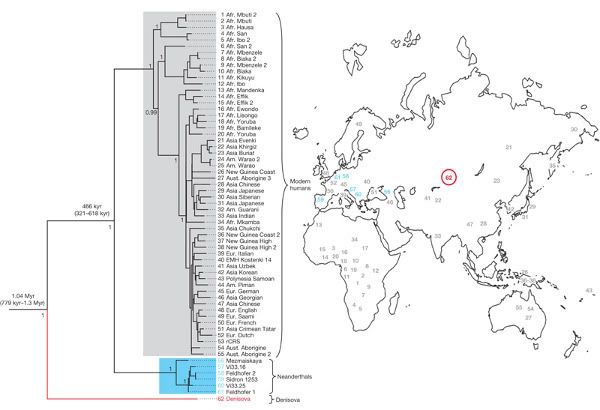
Here's the big figure from the paper, which was presented by Johannes Krause and colleagues in Nature yesterday. It's a phylogenetic tree which relates the little finger's mtDNA to H. sapiens and H. neanderthalensis sequences (click to see a high-resolution version):
The Denisnova sequence is red, Neanderthal sequences are in blue and modern humans are grey. So, the Denisova mtDNA forms a distinct lineage that isn't represented in modern humans or in previously published Neanderthal sequences. By using the tree as the basis for molecular dating the researchers were able to estimate that Denisova lineage separated from other human mitochondrial lineages between 0.78 and 1.3 million years ago. The temporal context the molecular dating adds to the phylogenetic tree helps to us understand where this new mitochondrial lineage might fit into humanity's family tree.
I've said before that most of our species' history was played out in Africa, and, in fact, the same is true when we step up a taxonomic level and look at our genus. All the human species that have been found outside of Africa descend from migrants that moved out of that continent at some stage. Here's a schematic representing some of the species in the wider human family tree and the timing of the migrations that moved them out of Africa.
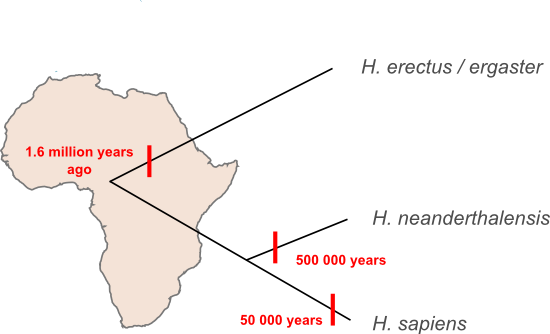
How does the new evidence presented by Krausse et al. fit into that scheme? Perhaps the simplest interpretation is the the Denisova lineage represents a new species. The estimated age of the Denisova lineage makes it too young to have been carried out of Africa by the first wave of H. erectus migrants to leave Africa and apparently too old to have been inherited from the migrants that went on to form the Neanderthal lineage. If the Denisova sequence is something new then we'll have to update our family tree, adding a new branch and a fourth migration out of Africa.
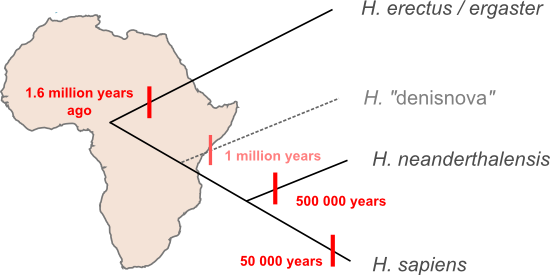
John Hawks thinks we should hold off on updating the family tree too qucikly. The Desinova specimen might be a Neanderthal. At first glance the tree presented by Krausse et al. seems to dispel that possibility since previously identified Neanderthal sequences are more closely related to modern human sequences than the new linaeage, but that tree is based entirely on mtDNA. The mitochondrial genome is inherited as if it was a single gene. We can often use trees estimated from a single gene ("gene trees") as a proxy for species-level relationships ("species trees") but, in fact, every gene in a population has its own history and there there are scenarios that can push a given gene tree away from underlying species tree. Perhaps the easiest way to visualise how you'd end up with mitochondrial lineages that diverged millions of years ago within a single species is to think about genetic lineages moving through a population while speciation happens. New species form when populations stop sharing genes with each other, in the diagram below the big black triangle represents a barrier to gene flow. What happens if multiple different gene lineages are present in the ancestral population at the time that this gene flow stops? Usually, given enough time, each species will "sort" into specific gene lineages that descend from just one of the lineages in the ancestral population, but it's also possible for one (or both) species to maintain multiple lineages for some time. Such "incomplete lineage sorting" makes gene trees bad proxies for species trees and it's just possible that something like this has happened in Neanderthals:
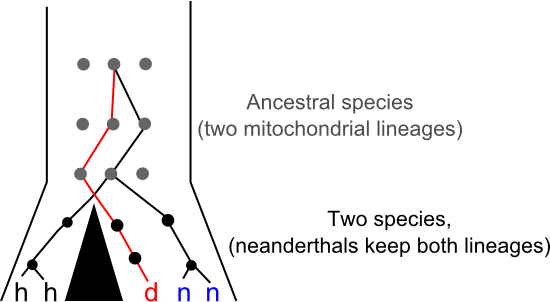
Perhaps by moving to the very Easterm edge of the Neanderthals range we've sampled for the first time a lineage that existed in that species for the whole time it was in Europe. Maybe, and Hawks surely knows a lot more about paleobiology than I do, but I don't really buy it. It's certainly possible for a species to harbour deeply divergent mitochondrial lineages, but the time it takes for gene-lineages to sort within a species is relative to the effect population size of that species. Neanderthals probably had a relatively small effective population size (and mtDNA definitely does, since only females pass it on and then in only one copy) making the retention of multiple lineages over hundreds of thousands of years seem like a long shot. As Hawkes argues, strong geographic structure in Neanderthal populations might have aided the retention of divergent genetic lineages against those odds, maybe the Denisova mitochondrial lineage was extinct in Western Europe but common in Central Asia? It's possible, but I wouldn't bet on it.
Finally, the Denisova sample might be our first look at H. erectus DNA. H. erectus remains have been recovered from China so it seems possible they were in Siberia too. As I've said, the molecular dating of the Denisova lineage probably makes it too young to be a descendant of the first wave of migration form Afirca (though, of course, there is some uncertainty associated with that dating), but it might be evidence of genetic exchange between African and the H. erectus diaspora. As we've come to understand the origin of our species we've realised that the simple "Out of Africa" model is just that, a model, and the true pattern is more complex. H. sapiens really did have its start in Africa and it really did push out into the rest of the world in the last 50 000 years or so, but during that expansion populations have continued to exchange genes. There's no reason to believe that that H. erectus could not have done the same, perhaps the main thrust of the H. erectus expansion was 1.6-2 million years ago but genes continued to flow in and out of Africa for sometime after that.
So, there are three possibilities for the Denisova sample:
- It could be a new species,
- It could be an ancient mitochondrial lineage retained in eastern Neanderthal populations but lost elsewhere
- It could be the first H. erectus sequence.
Krause, J., Fu, Q., Good, J., Viola, B., Shunkov, M., Derevianko, A., & Pääbo, S. (2010). The complete mitochondrial DNA genome of an unknown hominin from southern Siberia Nature DOI: 10.1038/nature08976
Labels: evolution, genetics, Human evolution, might interest someone, phylogenetics, sci-blogs, science, speciation
22 Comments:
A good question, and even the supplementary data to the Nature paper doesn't make it clear.
I have a feeling that you could design primers that are 'universal' enough to capture human mtDNA and specific enough to prevent you getting a lot of non target sequence.
But yeah, that don't spell it out for us
Genetic phylogeny is generally correct (and, of course, I assume that the aDNA sequencing is correct) but that only tells us that the Denisova hominin's mtDNA divergence time is, very roughly, double than Neanderthal's.
There's really no particular reason to believe that Neanderthals only diverged c. 700-300 Ka ago (the dates used in the paper). In fact something like 900 Ka is probably closer to reality, as those are the oldest known dates for Acheulean out of Africa, more exactly in Europe (where we also find all the line leading to neanderthals: H. antecessor, H. heidelbergensis and H. neanderthalensis itself).
Unless one postulates a back-migration to Africa of H. heidelbergensis leading to H. sapiens (what has no archaeological backing), that's the most reasonable estimate for the divergence between Neanderthals and us.
If the Neanderthal divergence date is of c. 1 million years, then the Denisova divergence date is of c. 2 million years, what makes her lineage part of the (archaeologically confirmed) H. erectus migration around that date. It might still have crossed into a Neanderthal population somehow or it might be a real surviving H. erectus, maybe of an evolved oriental branch like the Dali specimen.
Paleoanthropology isn't my thing, so I can't really argue with you about neanderthal divergence date. (I've read people that place H. rhodensis in with H, heidelbergensis and H. antecessor as an erectus derivative which would bring us back to 500k or so. But I don't have any reason to pick on scenario over another.)
As you say, there are assumptions underlying an molecular clock estimate and we definately shouldn't, like most of the media has, take the point estimate as the time of divergence. Having said that, I'd be surprised if a clock calibrated from the human-chimp split the sapiens-neanderthal split wrong by a factor of two.
It's definitely going to be interesting when the nuclear DNA comes out (or if they find a real complete specimen)
The Pan-Homo divergence date is also matter of controversy. I usually read dates ranging from 5 to 7 million years, however Caswell 2008 suggests at least 8 million years and, following her logic (Congo river formation directly related to Chimp-Bonobo split), could be as much as 10 million.
The lower dates for Pan-Homo split are based on what seems to be (I'm no expert either) a very limited fossil record, which can only provide minimal dates, not absolute ones.
Also other papers have recently demanded longer chronologies for the MC: Heads 2010 had to push the MC estimates to their maximum to allow for new world monkeys not to cross the ocean in a most improbable swimming feat. Roach 2010 found that the actual mutation rate in humans is a mere 40% the usual estimates.
The issue is unclear but I find that there are good reasons to suspect that the MC needs a lot of refining yet (probably towards longer chronologies) before it can produce reliable results we can trust at the same level as the archaeological record.
"It's definitely going to be interesting when the nuclear DNA comes out (or if they find a real complete specimen)".
Absolutely.
Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany. Svante Pääbo, director of the institute's genetics department, leads the Neanderthal Genome Project
some of their research indicates a genetic signal from H. Neander 1=4% present in all non-African humans
if true it might be fun to mention to the Aryan supremacist that they are not "pure strain" humans at all, only Black Africans are.
but I will wait for the next publication
It's already out! Looks pretty convincing to me (it's _just_ conceivable that we are actually looking at geographic variation in african genomes that we haven't yet sampled)
I'm planning to write something about it soon, but a bit snowed under at the moment. My sciblogs "scibling" Alison has a nice post here.
"The pinkie wa sfound in a 40 kyp level".
Not by this paper, which attributes the finger to >50 Ka ("infinite") radiocarbon dates, which are all the dates of pre-AMH occupations be them Neanderthal or "Denisovan" in the area. AMH occupations are from at least 48 Ka. on (further south) or 30 Ka on in Denisova cave itself.
Levels 20 to 12 are Middle Paleolithic, while Level 11 belongs to the Upper Paleolithic. Level 11 yielded bone implements,, personal adornments made of stone, bone, ostrich eggshell, mammoth ivory and animal teeth. In short the complete AMH package. The Levallois used in this region came from the Levant and was not made by Neandertals. Derevianko is quite clear on that.
That is not to say that AMHs and Neandertals didn’t share the region, which they obviously did. We need more than one just one pinkie to claim a new race of hominins.
sv77
depression treatment. Your weblog offered us priceless information to work on. You may have completed a marvellous job!
---------------------------------------------------------------
[url=http://www.sexybags.info/rssrock.html]My designer handabgs[/url]